The BigMac dataset - ultra-high angular resolution diffusion imaging and multi-contrast microscopy of a whole macaque brain
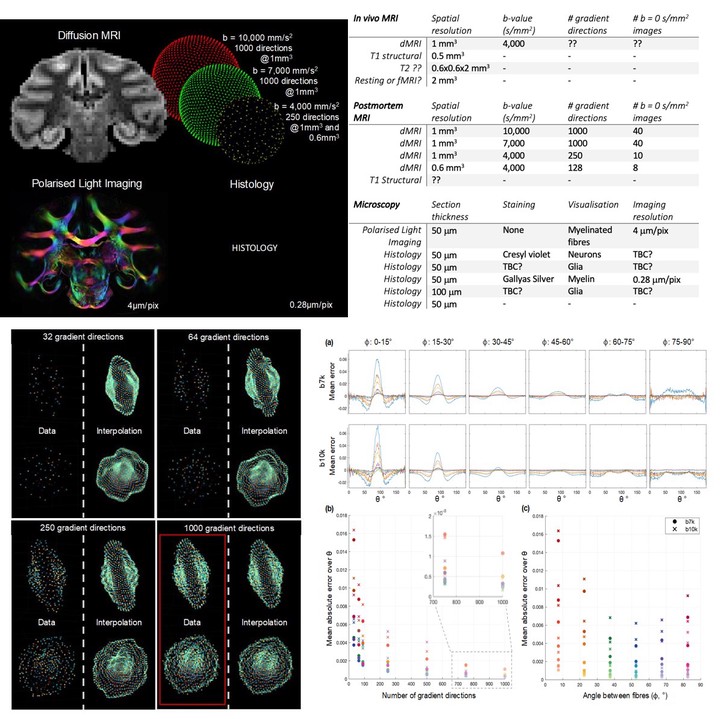 Image credit: From Abstract (Amy Howard)
Image credit: From Abstract (Amy Howard)Introduction:
Diffusion MRI (dMRI) has great potential for studying the complexity of white matter fibre architecture non-invasively. However, because dMRI is an indirect measure of this microstructure, we require validation datasets for two main purposes: (i) to relate dMRI to microscopy data that directly measures the microstructure of interest; and (ii) to relate high-quality dMRI data to more conventional data quality.
We present the NAMETBC dataset which addresses both of these goals. NAMETBC combines ultra-high angular resolution diffusion MRI (HARDI) at multiple b-values and resolutions with histology and polarised light imaging (PLI) data in a postmortem macaque brain. The same animal was also scanned in-vivo with dMRI. This dMRI dataset will enable us to characterise the dMRI signal in great detail. In particular, with 1000 gradient directions in the outer two shells, we retain dense sampling of the diffusion signal on any arbitrary 2D plane (Fig 2), aiding robust dMRI-microscopy data comparison. The NAMETBC dataset thus provides a platform from which we can interconnect microstructural features with dMRI signals throughout the entire brain.
To demonstrate one potential use of the NAMETBC dataset, we investigate to what extent the under-sampling of q-space biases our estimation of the ‘true’ diffusion signal of crossing fibres.
Methods:
An overview of the post-mortem acquisition of the NAMETBC data is given in Figure 1. dMRI data was acquired at 7T (SCANNER SPEC) with a total of 2500 gradient directions, b=4,000, 7,000 and 10,000 s/mm2 and two spatial resolutions (0.6 and 1 mm3) (Table 1). The adult macaque brain (male, 11.7years old) was then sectioned along the coronal plane at 50/100m thickness. Consecutive slices were processed for either histological staining, with the first three sections stained for neurons (Cresyl violet staining), glia (IBA-1, GFAP), and myelin (Gallyas silver staining), followed by an unstained slice for polarised light imaging.
To demonstrate a first application of NAMETBC, we consider the effect of the number of sampling directions on the estimation of the true underlying diffusion profile, as reflected in the full 1000 directions. The dMRI data (b = 7,000 and 10,000 s/mm2) was divided into subsets of 32, 64, 90, 250, 500 and 750 gradient directions whilst retaining uniform coverage of q-space [ref Camino,orderpoints]. Using distance-based interpolation, the diffusion profile of each subset was estimated along the original 1000 directions and compared to the fully-sampled data to generate an error. Here the ‘smoothness’ of the interpolation was related to the minimum solid angle between adjacent q-space samples. Figure 2 demonstrates the interpolation of a single voxel in the white matter. To study the effect of sampling on different fibre configurations, we fit the ball-and-stick model [ref] to identify one- and two-fibre voxels, then further categorised two-fibre voxels according to the incident angle between the fibres, . Finally, the error between the measured and interpolated signal was calculated as a function , the angle between the mean fibre orientation and the diffusion gradient.
Results:
Figure 4a shows how under-sampled data routinely fails to faithfully reconstruct the ‘true’ diffusion profile, often under- or over-estimating the diffusion signal perpendicular (=90) or parallel to the fibre respectively. Figures 4b,c summarises these plots as the mean across of the absolute errors for each category of . The mean absolute error varies approximately exponential with the angular resolution (4b). For highly sampled data, b = 10,000 s/mm2 begins to outperform b = 7,000 s/mm2 (see inset), suggesting that contrast is more important than SNR in this regime.
Figure 4c shows the error as a function of angle between fibres, suggesting a more accurate reconstruction of the diffusion profile of fibres crossing = 45-60. The high error at small demonstrates that low angular resolution data fails to capture important information in the diffusion signal that likely discerns multiple, highly-aligned fibres from disperse single fibre populations. This dependence of the error is less marked for high angular resolution data.
Discussion/Conclusion:
This study constitutes the first investigation of the NAMETBC dataset. These first results demonstrate that the accuracy of the diffusion profile increases exponentially with the angular resolution of sampling, and that high angular resolution is necessary to characterise acutely crossing fibres. As our analysis is independent of any particular diffusion model (i.e. is based purely on an interpolation of the signal to the locations of the 1000 direction data), these results have implications for all HARDI-based analyses.
This study has considered the highly-oversampled dMRI data of NAMETBC to represent the ground truth of the diffusion profile. Future work will focus on the harmonisation of the dMRI, histology and PLI data to both validate and drive biophysical modelling of the white matter microstructure.