Exploring the Proteome of ALS Laser Microdissected Purkinje Cells – Method Development.
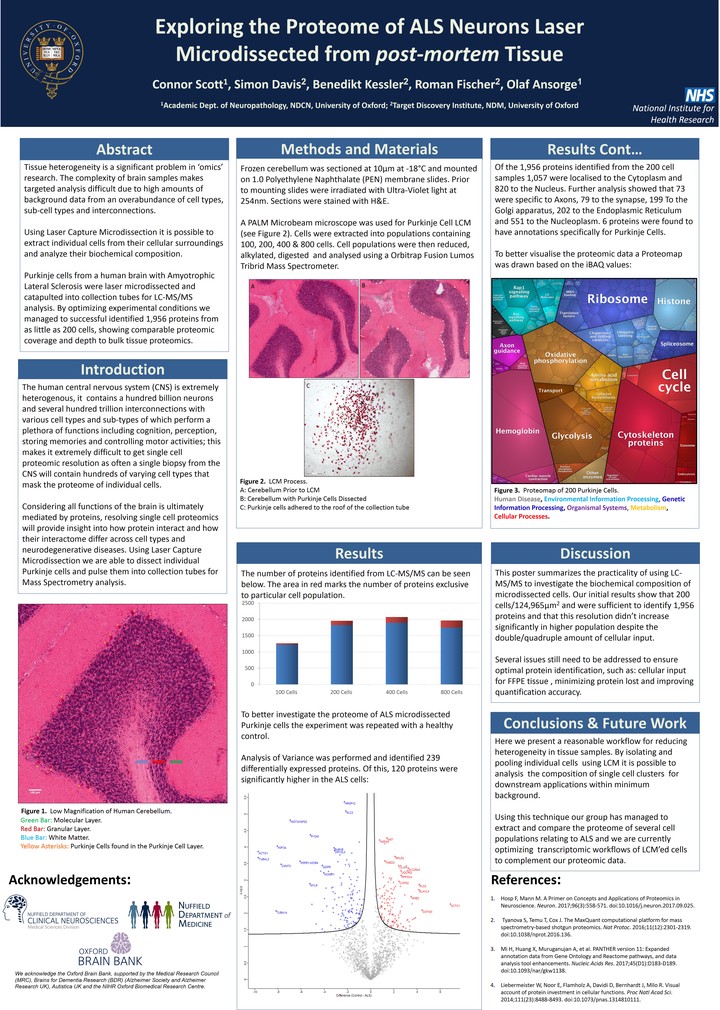 Image credit: Own Work
Image credit: Own WorkLiquid Chromatography-Mass Spectrometry (LC-MS/MS) sensitivity has improved significantly over the last decade and despite these developments, it has had a low impact in the field of neuroscience compared to other techniques such as MRI, in-situ hybridisation and optogenetics. This is partially due to the heterogeneous nature of the human brain which makes acquiring proteomic data from single cell populations challenging. The human central nervous system is extremely heterogeneous, it contains 100 billion neurons with several 100 trillion interconnections; collectively they perform a plethora of functions including cognition, sensory and motor control. All functions of the brain are ultimately mediated by proteins and exploring the proteome of individual cells can reveal additional functions and responds to neurodegenerative diseases. However, it is often difficult to extract single-cell proteomic data from the CNS as a single biopsy can contain hundreds of cell types which can mask individual proteomes. A targeted single-cell proteomic approach needs to be utilised to explore the pathology of individual cells. Our group has developed a ‘microproteomic’ method using Laser Capture Microdissection (LCM) to isolate single cell populations and analyse their proteins with LC-MS/MS. We have developed a workflow optimising sample dissection, histology, LCM, sample preparation, protein reduction and alkylation and LC-MS/MS analysis through our collaboration with the Target Discovery Institute. Using this approach we have increased the proteomic coverage and depth of our samples to a resolution comparable to bulk tissue proteomics, using an input of only 200 cells. We have obtained a dataset of ~2,500 proteins from individually microdissected Purkinje Cells from a post-mortem brain with Amyotrophic Lateral Sclerosis. The proteome of these Purkinje cells was analysed against control tissue. We found an enrichment of proteins relating to mRNA processing (p 1.29E-03), mRNA splicing (p 3.85E-02) and an increase in RNA Polymerase II Transcription (p 9.53E-03) and RNA Polymerase II Regulation (p 1.71E-02). We also see a 6-fold enrichment in proteins that epigenetically regulate gene expression (p 3.20E-03). Our microproteomic approach is now being used to explore the differing proteomes of individual cells in ALS. This technique allows cells that were previously thought too difficult to remove from their cellular environment to be researched with higher sensitivity.